The Focinator v2
A new open-source tool for automated high-throughput foci counting
Focinator is an open-source software tool providing automated analysis of foci images including foci counting and colocalization of foci.
Try Focinator v2-40beta and let us know how you like it!
- We fixed an issue with "Remove Scale". The beta will only produce results in pixels
- We are working on the implementation of a “Remove Scale” option for e.g. area measurements
Out now: Focinator v2-31
-- New intensity measurements added for total nucleus and foci! --
Colocalization in two directions; your own channel names; Noise, cutoff, area, etc. values in the report file of your foci counting run.
Including new full automatic 4-channel analysis with up to 3 foci channel, new graphical user interface and colocalization! Try the new version now and help us developing the Focinator v2.
Please follow us on ResearchGate for latest news and patch notes!
researchgate.net/project/The-Focinator
Citing the Focinator!
Dear user of the Focinator,
We are pleased that the Focinator has awaken so much interest and is now used by many groups around the world. We would be very grateful if you could cite us in return:
Oeck et al. (2017) The Focinator v2-0 - Graphical interface, four channels, colocalization analysis and cell phase identification. Radiat Res 11. PMID: 28492345
Oeck et al. (2015) The Focinator - a new open-source tool for high-throughput foci evaluation of DNA damage. Radiat Oncol 10(163):015-0453. PMID: 26238507
Using the Focinator!
The Focinator v2 is based on an ImageJ macro and an R script. To provide easy, user friendly image handling we use the Bio-Formats ImageJ plugin to guarantee, that most of the different image types will run with the Focinator v2. The latest version of the Focinator (v2-31) is now available on the download page. Follow the instructions to install and launch the Focinator. The Focinator v2 opens now automatically all images in a selected folder set and analyses all multi-channel foci images, identify the nuclei and measure their parameters. In the next step the Focinator v2 uses the marked nuclei as "Regions Of Interest" (ROIs) and evaluates the foci in one, two or three foci channel at the same time. Refer to the manual to learn using the Focinator v2.
Highlights!
- High throughput automated foci counting: faster and less prone to human bias
- Robust method for recognizing cells or nuclei as ROIs
- Measures area, min, max and mean intensity of each ROI and up to three foci channels
- Offers single value proof and direct mean export
- Live counting of foci for reliable checking
- Evaluation of colocalization based on counted foci
- Data collection and further processing in Excel of your foci count
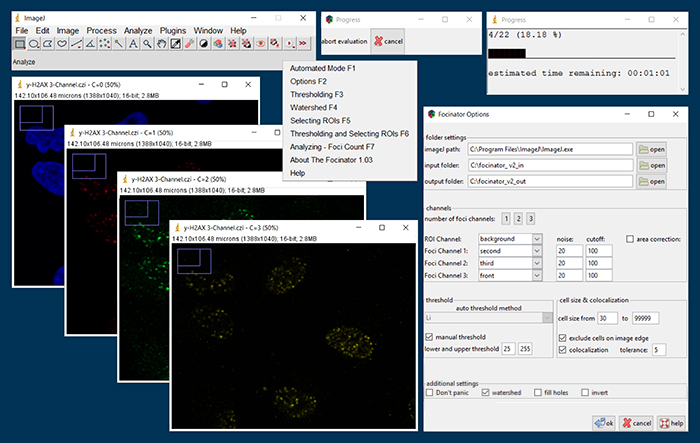
The new interface of the Focinator v2: